I've been brushing up on the ABC's of flower formation, and I want to see if I understand it well enough at this point to explain it to anyone else. Readers of this blog are therefore going to be my guinea pigs.
I'm using mostly the names of the genes as they were described for the dwarf cress, Arabidopsis thaliana, which is the "lab rat" or "fruit fly" for plant biologists. Arabidopsis thaliana is a small species, easy to propagate in the lab, with a few large chromosomes, and one from which it has been possible to isolate a large number of mutants.
Almost 20 years ago, it was found that there are a set of specific genes that lead a new shoot to form a flower. Because there were initially three general types found, they were referred to as the A, B, and C classes. If you inactivated certain of the genes, you did not get any petals in the flower. This was therefor called APETALA. When it was found that there were two genes that caused loss of petals when mutated, they ended up with APETALA1 (AP1) and APETALA3 (AP3). It turns out that you need two classes of genes to make petals: A and also B. APETALA1 is the A gene and APETALA3 is one of the B genes. APETALA probably comes from "a-" meaning without and "petal." A second B gene was found, and given the name PISTILLATA (PI). I haven't tracked down where that name came from. It turns out that there have to be functioning copies of both the B genes in order to have full, normal flower development. APETALA3 and PISTILLATA form what is called a "heterodimer," e.g., a structure made up of two protein molecules that are not identical.
The C gene is necessary for formation of the sexual parts of the flower. When it is mutated, there are no stamens and no pistil, so it was named AGAMOUS (AG). If one of the B genes is mutated, you get the pistil but no stamens. The formation of stamens therefor requires functional B genes and the C gene. The C gene alone is needed for formation of the pistil.
When the A, B, and C genes are expressed ectopically (in non-flower parts of the plant), there are no flower parts formed. This indicated that more genes are needed to get sepals, petals, stamens, or pistils than just the ABC set.
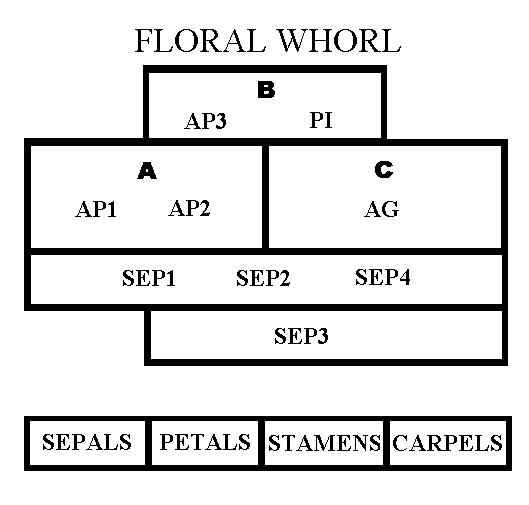
The Scheme for Floral Gene Interactions
Those additional genes are now referred to as the SEPALLATA (SEP) genes, and are often called the E class of genes. There are 4 of these genes, SEP1, SEP2, SEP3, and SEP4. These function somewhat redundantly, but in the absence of all 4 of the SEP genes, there are no flower parts produced. Which SEP genes are expressed depends on when and where one looks. The discovery that a fourth class of genes is required has led to recent papers calling this the "ABCE" model.
The "quartet model" for how these gene product work is based on studies of protein-protein interactions. The quartet AP1-AP1-SEP-SEP leads to formation of sepals. The quartet AP1-AP3-PI-SEP leads to petals; AP3-PI-AG-SEP induces stamens; and AG-AG-SEP-SEP forms carpels (ovary and pistil). Protein chemists would refer to these quartets as tetramers, since the four separate protein molecules involved are physically associated with each other.
All of these genes are members of the MADS-box family (except for AP2). The MADS structure is named for the four very different species in which such genes were found: Mouse, Arabidopsis, Drosophila (fruit fly), and Saccharomyces (yeast). MADS genes have also been found in mammals, including humans. This is clearly a very old motif in the history of life.
A paper published in 2007 on this is Soltis et al., The ABC Model and its Applicability to Basal Angiosperms, in ANNALS OF BOTANY, vol. 100, pp. 155�163. I also found a slightly older paper, Zahn et al., The Evolution of the SEPALLATA Subfamily of MADS-Box Genes..., in GENETICS, vol. 169, pp. 2209-2223 (2005) to be quite useful. If you search in Google Scholar using advanced search mode and including only recent years, you can find a large number of recent papers on the genetics of floral identity and determination. Try using search words like ABC + Floral, or MADS-box, or SEPALLATA, for instance.
Good gardening,
Jim